DNA
DNA4mC-LIP: a linear integration method to identify N4-methylcytosine site in multiple species
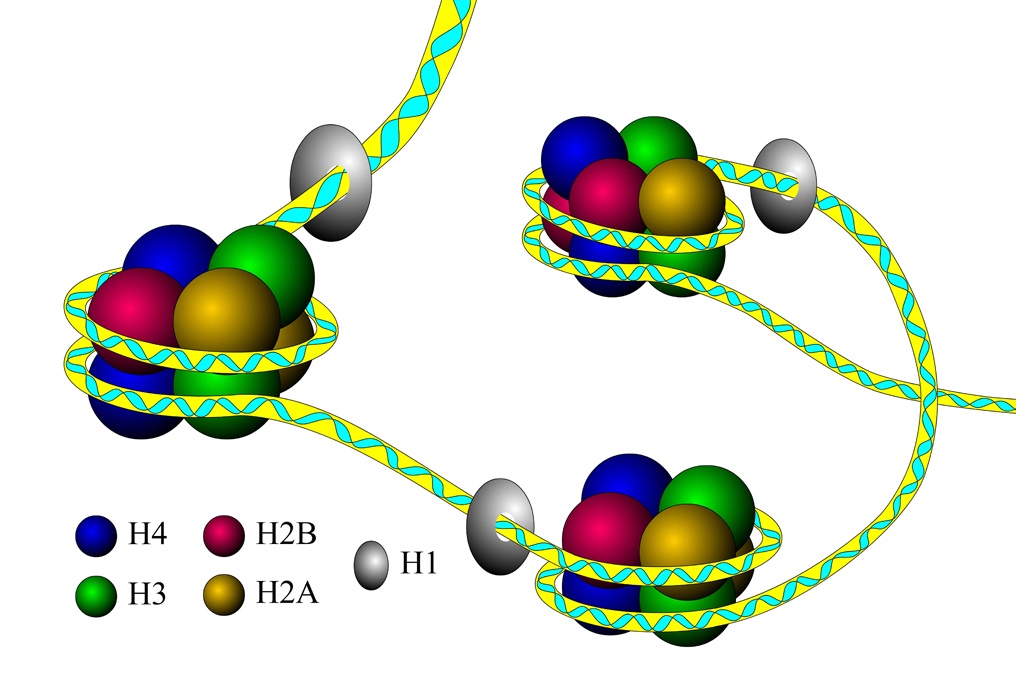
iNuc-PhysChem: Identification of nucleosomes in S. cerevisiae genome
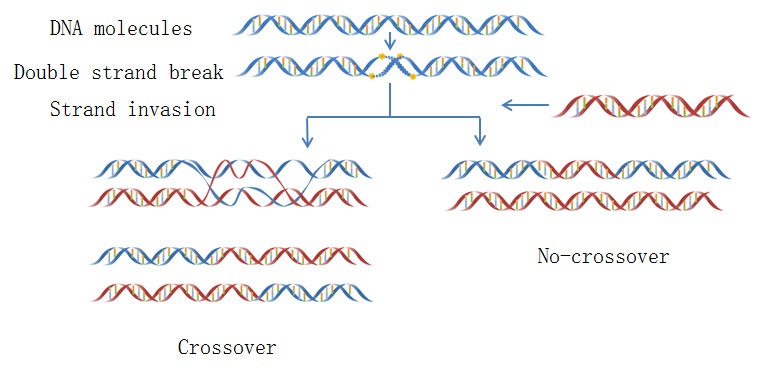
iRSpot-PseDNC
,
iRSpot-Pse6NC
and
iRSpot-Pse6NC2.0
: Prediction of recombination spots in S. cerevisiae genome
iNuc-PseKNC: APredicting nucleosomes in H. sapiens, C. elegan, D. melanogaster genomes
iTIS-PseTNC: Identification of translation initiation site in human genes
iPro54-PseKNC
and
iPro70-PseZNC
: Identification of sigma54 and sigma70 promoter in prokaryotic genomes
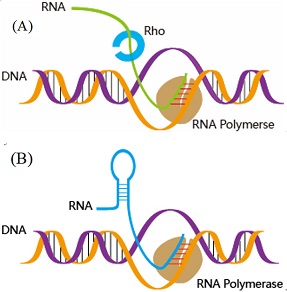
iTerm-PseKNC: Identifying bacterial terminators
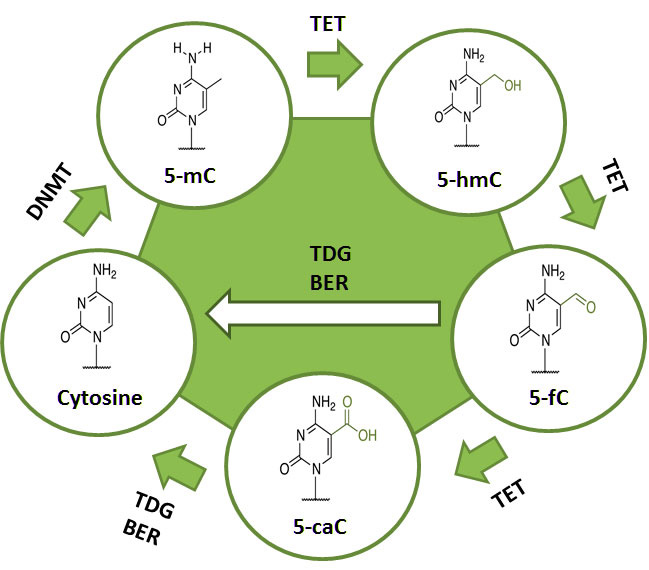
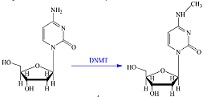
iDNA4mC: identifying DNA 4-methylcytosine sites
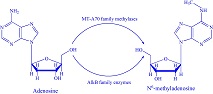
iDNA6mA-PseKNC
and
i6mA-Pred
:Identifying DNA N6-methyladenosine sites
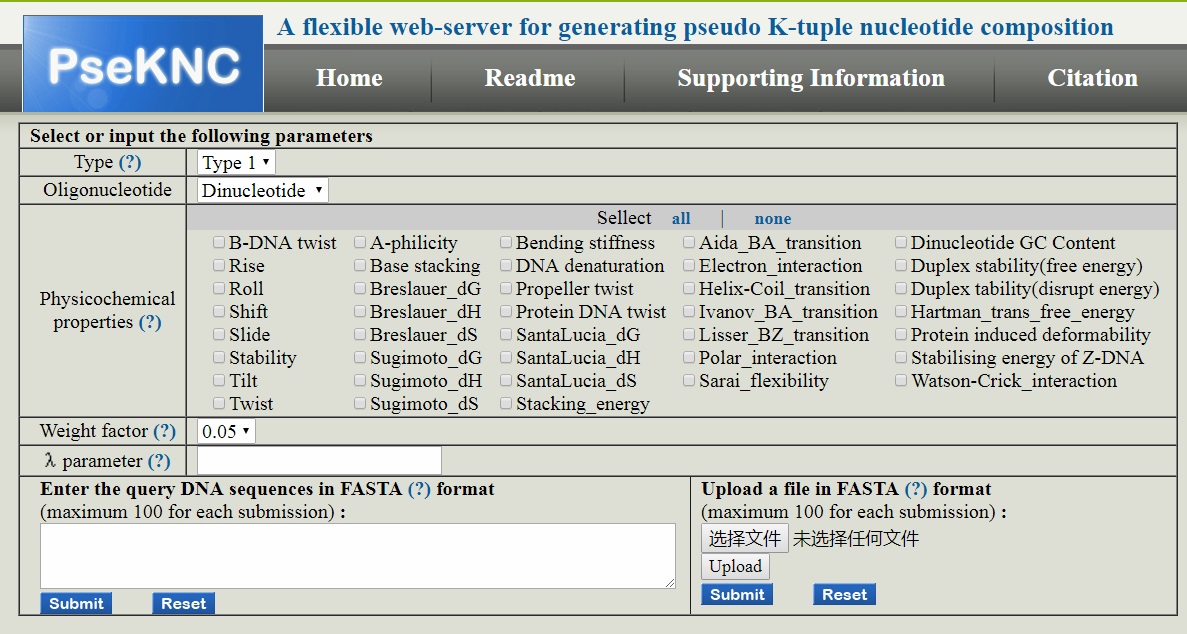
PseKNC
and
PseKNC-General